Executive Summary
Hyrax Biosciences’ Exatype platform provides automated, cloud-based analysis for accurate Mycobacterium tuberculosis (Mtb) drug-susceptibility testing (DST) and lineage typing from next-generation sequencing (NGS) data.
Validated using Agilent’s SureSelect XT CD Mtb panel, supplied by Diagnostech, the Exatype solution demonstrated high analytical accuracy, reproducibility, and compliance with WHO reference standards. Seamlessly integrating with existing sequencing workflows, Exatype converts raw data into high-confidence genomic results and interpretive reports - a rapid, scalable, and insight-driven solution that reduces the need for in-house bioinformatics and supports deployment across research and public health settings.
Key Outcomes:
- Specificity and Sensitivity: Complete (100%) concordance with known DST results from published data. Detection threshold: ≥10% variant prevalence detection at ≥30x coverage.
- Lineage classification: 100% detection across all samples using dual SNP-barcoding algorithms, offering enhanced resolution and confirmatory accuracy, compared to other tools.
- Conformance to the WHO Mutation Catalogue1: Exatype meets WHO v2 reference standards for mutation interpretation, combining transparent coverage reporting, proven analytical performance, and robust lineage typing through dual SNP-barcoding (Fig. 1).
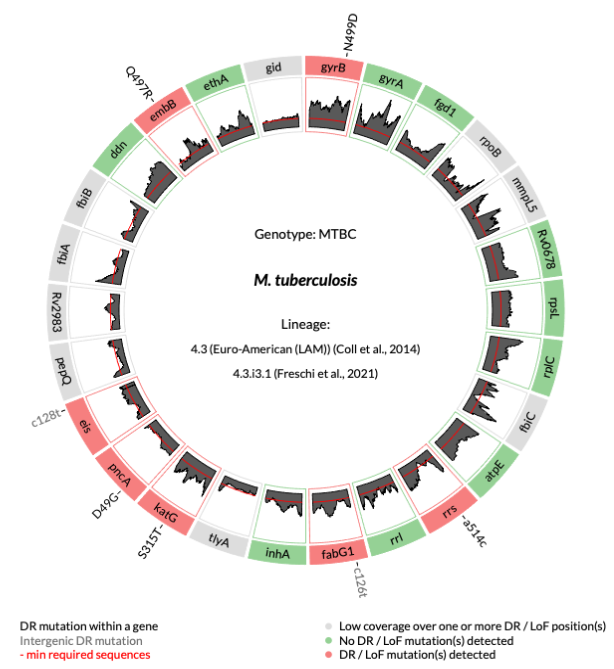
Background & Rationale
Tuberculosis (TB) affects over 10 million people annually, with ~3.3% of new cases being drug resistant. The World Health Organisation (WHO) recommends NGS for Mtb DST due to its accuracy, speed, and comprehensiveness relative to phenotypic testing (WHO consolidated guidelines on Tuberculosis2).
Agilent’s SureSelect XT CD Mtb Panels use precision hybrid-capture to enable DST from whole-genome (WGS) or targeted coverage directly from sputum, bypassing culture and reducing turnaround time. Hyrax Biosciences’ Exatype platform, incorporating both targeted and WGS Mtb pipelines, performs integrated quality control, drug-susceptibility reporting based on WHO-validated drug resistance associated mutations, and lineage typing using dual SNP-barcoding methods described by Coll et al., 20143 & Freschi et al., 20214. This study provides an analytical validation of Exatype’s performance on data generated using a Mtb whole-genome panel demonstrating a reliable, high-throughput solution for Mtb DST and genomic surveillance.
Validation Results
Study design: Fourteen Mtb samples (six from the original community design publication (Brown et al., 20155) and eight additional clinical isolates) were processed using the SureSelect XT CD Mtb Panel and analysed on the Exatype platform using the Exatype Mtb (WGS) product. Results were benchmarked against the expected outcomes described in Brown et al., 20155, and compared to three alternative Mtb pipelines.
Table 1: Drug-Susceptibility Reporting
| Metric | Exatype Result | Comparative Performance |
|---|---|---|
| Specificity | 100% concordance with known DST results* | Incorporating the updated WHO catalogue of mutations (version 2)1 enabled Exatype to report additional DST results, beyond what was reported by Brown et al., 20155 |
| Sensitivity threshold | Detects variants at ≥10% with ≥30× coverage | Confident detection of low-abundance mutations |
| Cross-platform concordance | ≥ 98% concordance across platforms | Equivalent or superior to other Mtb data-analysis pipelines |
*across drug resistance associated regions with sufficient coverage (≥30x).
Table 2: Lineage Typing
| Metric | Exatype Result | Notes |
|---|---|---|
| Specificity and Sensitivity | 100% concordance between SNP-barcoding algorithms | Dual SNP-barcoding enables cross-validated lineage calls - a feature offered by only 1 of the 3 other platforms. |
| Concordance with other platforms | Exatype matched or exceeded lineage calls from other tools | Confirms that Exatype delivers robust and reproducible classification |
Key Observations:
- Comprehensive Concordance: Full alignment with published resistance profiles for DR-, MDR-, pre-XDR-, and XDR-TB cases.
- Transparent Reporting: Low-coverage regions automatically flagged, maintaining interpretive integrity and traceability.
- Optimised Data Retention: Stringent QC processes preserves analytical depth while minimising artefacts.
- Enhanced Interpretation: Incorporation of the WHO v2 mutation catalogue extends the scope of detectable resistance mutations beyond prior studies.
Analytical and Applied Implications Enabled by Hyrax Biosciences’ Exatype Platform
This validation demonstrates Exatype’s seamless integration with established Mycobacterium tuberculosis sequencing workflows, converting raw data into reliable, high-confidence drug-susceptibility and lineage typing results.
For Hyrax Biosciences, this validation again highlights Exatype as a scalable, cloud-based analysis platform that standardises genomic interpretation in line with WHO-mutation catalogues. Integrated reporting, transparent quality control, and traceable results reduce the need for specialist bioinformatics and support regulatory and clinical adoption.
For partners, distributors, and assay developers, Exatype offers a ready-to-integrate analytics backbone that enables the delivery of end-to-end sequencing and interpretation solutions - accelerating implementation, expanding assay utility, and supporting data comparability across regions and technologies.
Conclusion
This validation demonstrates Exatype’s capacity to deliver consistent, high-quality analytical outcomes. By enabling standardised, transparent analysis aligned with WHO mutation catalogues, Exatype supports the global expansion of NGS-based TB research and surveillance.
With its adaptable architecture, Hyrax Biosciences is uniquely positioned to lead pathogen sequencing and analysis, empowering assay developers and advancing antimicrobial resistance genomics across global applications.
References
- World Health Organisation (WHO) Catalogue of mutations in Mycobacterium tuberculosis complex and their association with drug resistance, 2nd Ed
- WHO consolidated guidelines on tuberculosis: module 3: diagnosis: rapid diagnostics for tuberculosis detection, 3rd Ed
- Coll et al., 2014, A robust SNP barcode for typing Mycobacterium tuberculosis complex strains, Nature Communications, 5: 4812
- Freschi et al., 2021, Population structure, biogeography and transmissibility of Mycobacterium tuberculosis, Nature Communications, 12:6099
- Brown et al., 2015, Rapid Whole-Genome Sequencing of Mycobacterium tuberculosis Isolates Directly from Clinical Samples, Journal of Clinical Microbiology, 53(7):2230-2237
 Your sequence data.
Your sequence data.