Executive Summary
The COVID-19 pandemic underscored the global need for fast, accurate, and scalable pathogen identification. As respiratory-virus testing expands beyond single-target assays, the combination of multiplexed amplification panels and next-generation sequencing (NGS) has emerged as a powerful approach for comprehensive respiratory pathogen surveillance and identification. At the center of this evolution, Hyrax Biosciences’ cloud-based genomic analysis platform, Exatype, provides a platform-agnostic analysis environment capable of transforming raw sequencing data into actionable genomic insights.
In this analytical validation study, Exatype was used to analyse datasets generated from multiple respiratory virus panels across a range of sequencing platforms.
Exatype’s Key Capabilities:
- Identification of the infecting virus or co-infections with high confidence,
- Generation of consensus sequences, foundational for novel assay design and continuous lineage tracking,
- Analysis and visualisation of coverage across each target region to assess sequencing performance, and
- Quantification of amino acid and nucleotide variant prevalences, providing insight into potential changes in viral fitness, transmissibility, or treatment resistance.
Background & Rationale
The emergence of SARS-CoV-2 highlighted the limitations of traditional diagnostic approaches. During the COVID-19 pandemic, timely identification of pathogens - and their evolving variants - became critical to both clinical decision-making and public health response. Exatype played a pivotal role - processing approximately 42% of all SARS-CoV-2 genomes generated across the African continent (>72,000 genomes). This contribution established Hyrax Biosciences as a trusted analytical partner in large-scale genomic surveillance and rapid outbreak response.
Building on this foundation, we present an analytical validation study demonstrating Exatype’s capability for processing multi-target respiratory virus panels. These assays, which amplify a wide range of pathogens from a single sample, have transformed respiratory testing. When combined with NGS and an analysis solution like Exatype, these assays enable comprehensive and rapid identification of viruses such as SARS-CoV-2, Influenza A/B and Respiratory Syncytial Virus (RSV).
Validation Results
Study Design
Exatype was used to analyse raw sequencing data from twenty respiratory virus samples obtained from public (n=16)1, 2, 3 and collaborator (n=4) sources. The data was generated with different multi-target assays (including Illumina Respiratory Virus Oligo Panel (RVOP), Thermo Fisher Scientific’s Ion AmpliSeq FluAB Research Panel, and Twist Bioscience Respiratory Virus Panel) across Illumina, Ion Torrent, and Oxford Nanopore Technologies platforms, using standardised, platform-specific workflows.
Quantitative Outcomes
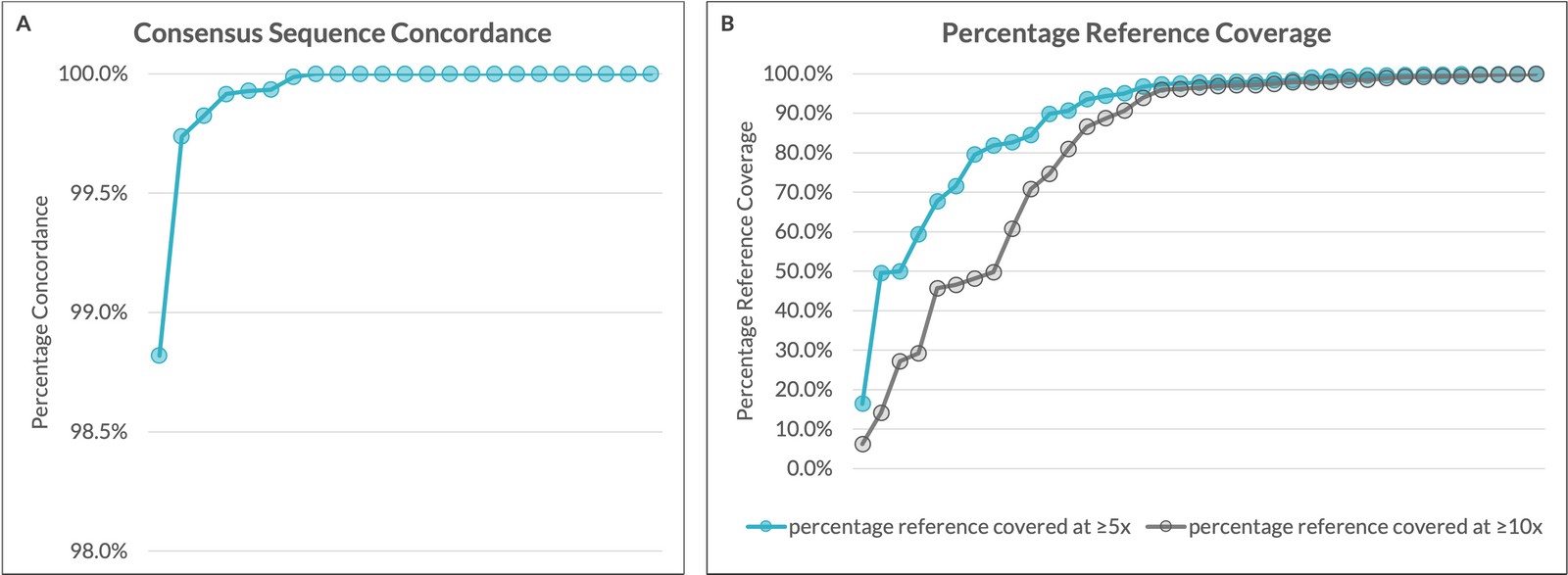
Table 1: Key Outcomes Summary
| Analytical Metric | Validation Result | Notes |
|---|---|---|
| Pathogen Identification | 100% match to expected respiratory viruses* | Accurate identification of all viruses present, including mixed infections |
| Consensus Sequences | ≥98.8% concordance*# (Fig. 1A) | High reproducibility confirms analytical robustness |
| Coverage Depth | ≥95% of reference covered at ≥5×; ≥85% covered at ≥10× across samples and platforms (Fig. 1B) | Coverage varied across samples and platforms but consistently met analytical thresholds |
| Sensitivity Threshold | Detection of minor variants ≥10% | Enables monitoring of within-sample diversity and emerging mutations |
*for samples and/or regions with sufficient coverage (≥20x for Illumina paired-end reads, ≥30x for Illumina single-end reads, Ion Torrent and ONT)
#where comparator consensus sequences were available
Exatype: Analytical Validation and Real-World Genomic Insight
Analytical Insights: Precision Through Integration
Exatype provides a comprehensive analytical framework for multi-target respiratory virus sequencing, delivering precision and depth beyond standard pathogen identification. Each analysis produces complete consensus sequences, coverage profiles across all target regions, and quantitative amino acid and nucleotide variant prevalences, all generated within a single, integrated workflow. These results enable laboratories and assay developers to evaluate assay performance, confirm amplification success, and monitor emerging mutations with implications for viral fitness, transmissibility, or therapeutic response. By consolidating these functions into one validated platform, Exatype ensures analytical consistency and reproducibility across sequencing technologies and assay formats.
Practical Value: From Analysis to Application
The analytical capabilities of Exatype extend beyond assay-specific performance, supporting standardisation, reproducibility, and transparent data interpretation across sequencing technologies. By integrating Exatype, laboratories and assay developers gain a unified framework for comparative analysis, performance benchmarking, and variant tracking. This interoperability strengthens data integrity, accelerates research translation, and supports surveillance initiatives where consistent, high-quality results are essential.
Conclusion
This analytical validation confirms that the Exatype platform provides accurate, reproducible, and biologically meaningful analysis of multi-target respiratory virus panels. Across three leading sequencing technologies and multiple commercial assays, Exatype consistently reproduced expected results with high concordance - while providing enhanced visibility into viral genome coverage and variation.
As respiratory virus testing evolves beyond simple detection, Exatype enables laboratories to extract deeper genomic insight from their data - supporting not only identification but also surveillance, variant tracking, and public health preparedness. In doing so, Exatype establishes an analytical standard for precision, scalability, and insight in pathogen genomics.
References
- Mu et al., 2023, The combined effect of oseltamivir and favipiravir on influenza A virus evolution in patients hospitalized with severe influenza, Antiviral Research, 216: 105657
- Goya et al., 2023, Genomic Characterization of Respiratory Syncytial Virus during 2022-23 Outbreak, Washington, USA, Emerging Infectious Diseases, 29 (4): 865-868
- Mehta et al., 2021, Respiratory Co-Infections: Modulators of SARS-CoV-2 Patients’ Clinical Sub-Phenotype, Frontiers in Microbiology; 12: 653399
 Your sequence data.
Your sequence data.