Executive Summary
Hyrax Biosciences, through its Exatype platform, delivers scalable HIV drug-resistance analysis to global programmes - processing over 150,000 HIV sequences for 1,146 users across 70+ organisations in 103 countries. More than half of these programmes operate in Global Health Equity settings, positioning Exatype as one of the most widely adopted and trusted platforms for routine Sanger-based HIV genotyping worldwide.
Building on this foundation, Hyrax Biosciences validated whether Illumina NGS data generated using the Thermo Fisher HIV-1 Genotyping Kit with Integrase, when analysed on Exatype, delivers drug-resistance results equivalent to the Sanger (capillary electrophoresis sequencing) gold standard. This analysis shows strong cross-platform concordance, demonstrating that the kit performs consistently whether sequenced on Illumina or Sanger. Exatype was used as the common analytical pipeline, ensuring standardised processing and fully comparable outputs.
Key Outcomes:
- Specificity: >98.7% concordance between Illumina and Sanger consensus sequences.
- Repeatability: ≥99.0% intra-sample concordance across replicates.
- Sensitivity: Equivalent detection of mutations at ≥20% prevalence (Sanger gold standard); Illumina additionally resolves variants below this threshold.
- Throughput: Illumina generated ~340,000 reads per replicate, delivering high coverage across PR, RT and IN targets.
Background and Rationale
At the end of 2022, 29.8 million people were on antiretroviral therapy (World Health Organisation, HIV Statistics, 2025)1, yet HIV drug resistance remains a major global health threat. Global HIV drug-resistance monitoring requires validated workflows that are accurate, scalable, and compatible across sequencing technologies. While Sanger sequencing remains widely used, laboratories are increasingly shifting toward NGS platforms such as Illumina to improve throughput and sensitivity.
Hyrax Biosciences undertook this analytical validation to assess whether Illumina sequencing data generated using the Thermo Fisher ABI HIV-1 Genotyping Kit with Integrase, analysed on the Exatype platform, provides results consistent with Sanger sequencing while enabling deeper variant profiling. Both the Sanger sequencing and Illumina NGS data were processed using Exatype ensuring directly comparable outputs.
Validation Results
Study design: included 5 samples × 4 replicates for PR (protease) and RT (reverse transcriptase) - sequenced as a single amplicon - and IN (integrase). All samples and replicates were amplified with the Thermo Fisher ABI HIV-1 Genotyping Kit with Integrase, sequenced on Illumina and Sanger, and analysed on Exatype.
Quantitative Outcomes
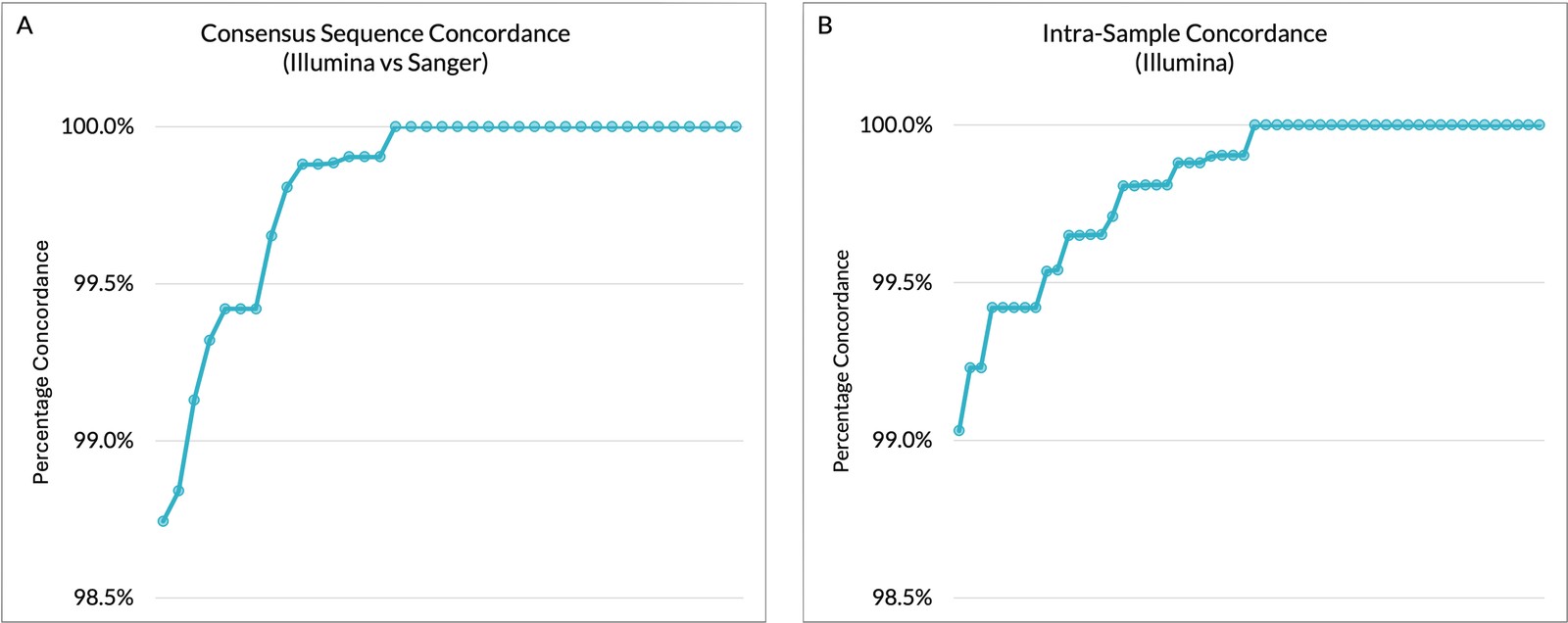
| Metric | Result | Benchmark / Notes |
|---|---|---|
| Specificity | >98.7% Illumina-Sanger consensus concordance (Fig. A) | Confirms platform equivalency |
| Repeatability | >99.0% concordance across replicates (Fig. B) | Demonstrates high robustness |
| Sensitivity | Matching detection of major variants (≥20% Sanger standard) | Equivalent performance to Sanger |
| Sensitivity threshold | Sanger threshold 15 - 20%2; Illumina threshold ≥1% | Enhanced minor-variant resolution with Illumina |
| Read depth | ~340,000 reads per replicate; ≥89% mapped | Supports high-confidence variant calls |
| Coverage | >15,000× average across PR, RT, IN | Exceeds minimum required (≥14x) for DR calling |
| Drug-resistance calls | Consistent across platforms | Reliable HIV-DR detection |
Key observations:
- Reproducibility across Illumina and Sanger sequences was ≥98%, confirming assay performance.
- Minor differences between Illumina and Sanger (and among replicates) were primarily attributable to shifts in variant prevalence at mixture sites.
Analytical and Applied Implications enabled by Hyrax Biosciences’ Exatype Platform
This validation confirms that programmes can continue using the same Thermo Fisher ABI HIV-1 Genotyping Kit they have relied on for years, while seamlessly expanding from Sanger to Illumina without changing their analytical workflow. By processing both Sanger and Illumina data through the same Exatype pipeline, laboratories retain identical interpretation rules, mixture thresholds, reporting formats, and drug-resistance outputs - ensuring continuity across longitudinal datasets.
Illumina’s high read depth, combined with Exatype’s automated QC and unified variant-calling framework, offers improved minor-variant detection without altering established Sanger-based thresholds. This cross-platform consistency allows laboratories to increase throughput and sensitivity using familiar reagents and a familiar analysis environment, removing the need for new bioinformatics tools or retraining. In practice, it reduces operational burden, supports scalable national surveillance, and keeps historical Sanger data comparable as programmes transition to NGS.
Conclusion
Illumina sequencing, generated using the Thermo Fisher ABI HIV-1 Genotyping Kit with Integrase and analysed through Hyrax Biosciences’ Exatype platform, provides HIV-1 drug-resistance results equivalent to Sanger while adding the benefits of high throughput and improved minor-variant resolution. This analytical validation confirms Exatype as a robust, cross-platform solution capable of supporting large-scale HIV-DR surveillance and laboratory implementation across diverse sequencing environments.
 Your sequence data.
Your sequence data.